Research
My doctoral research lies at the intersection of graph neural networks, computational biology, and large-scale data analytics. I focus on developing novel deep learning architectures for drug discovery and molecular property prediction, leveraging big data to accelerate scientific breakthroughs.
Publications
Papers and ongoing research work
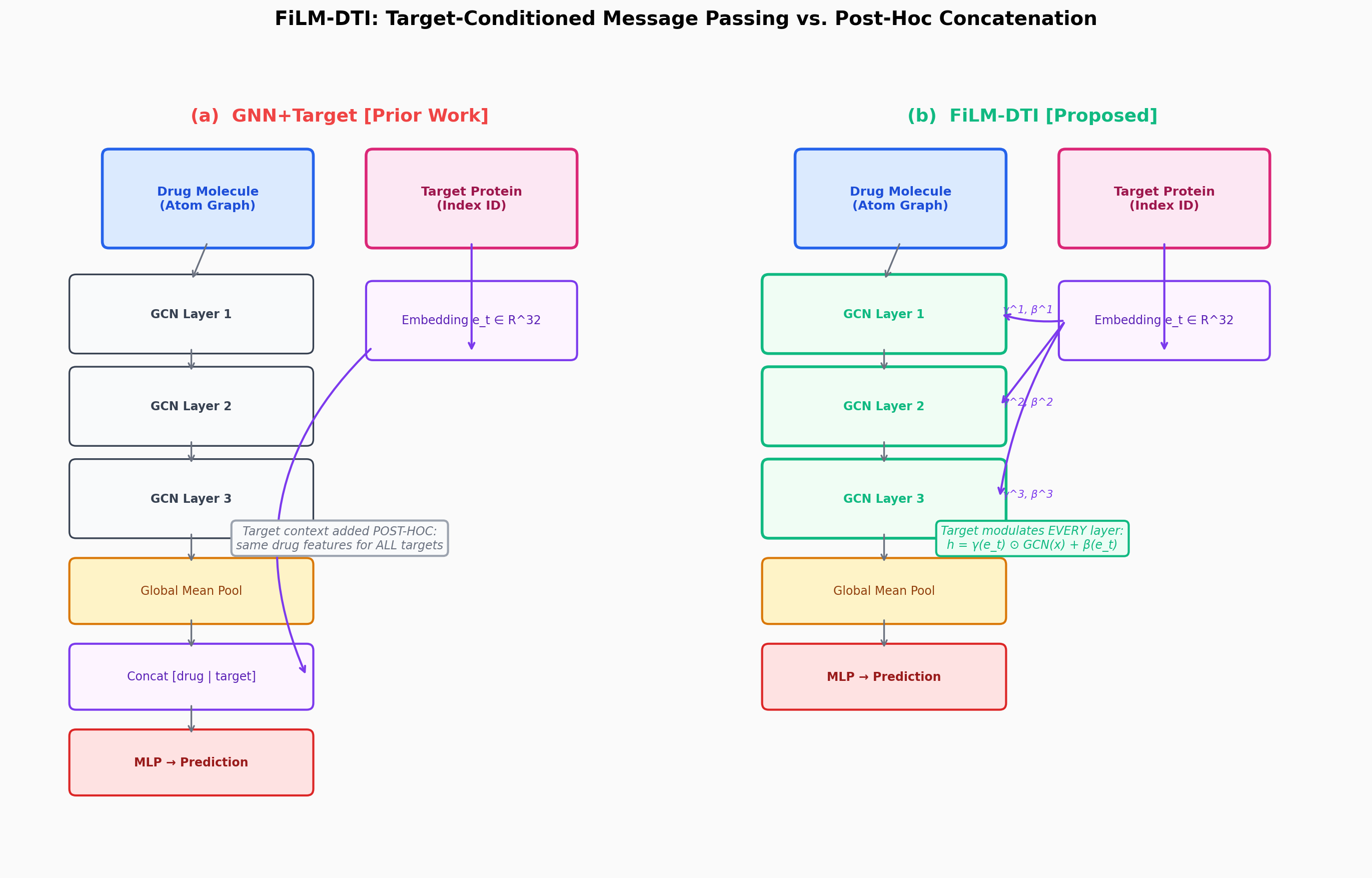
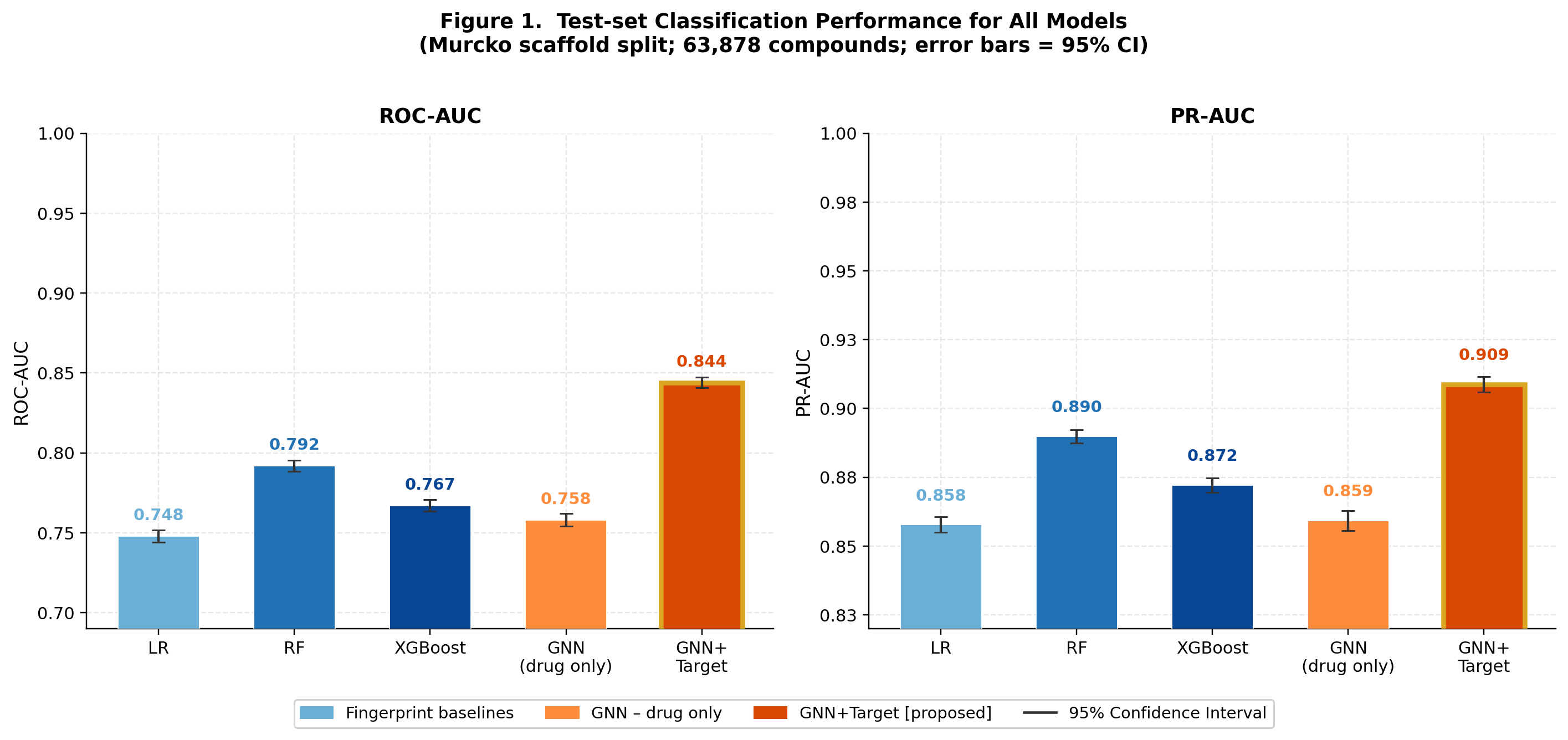
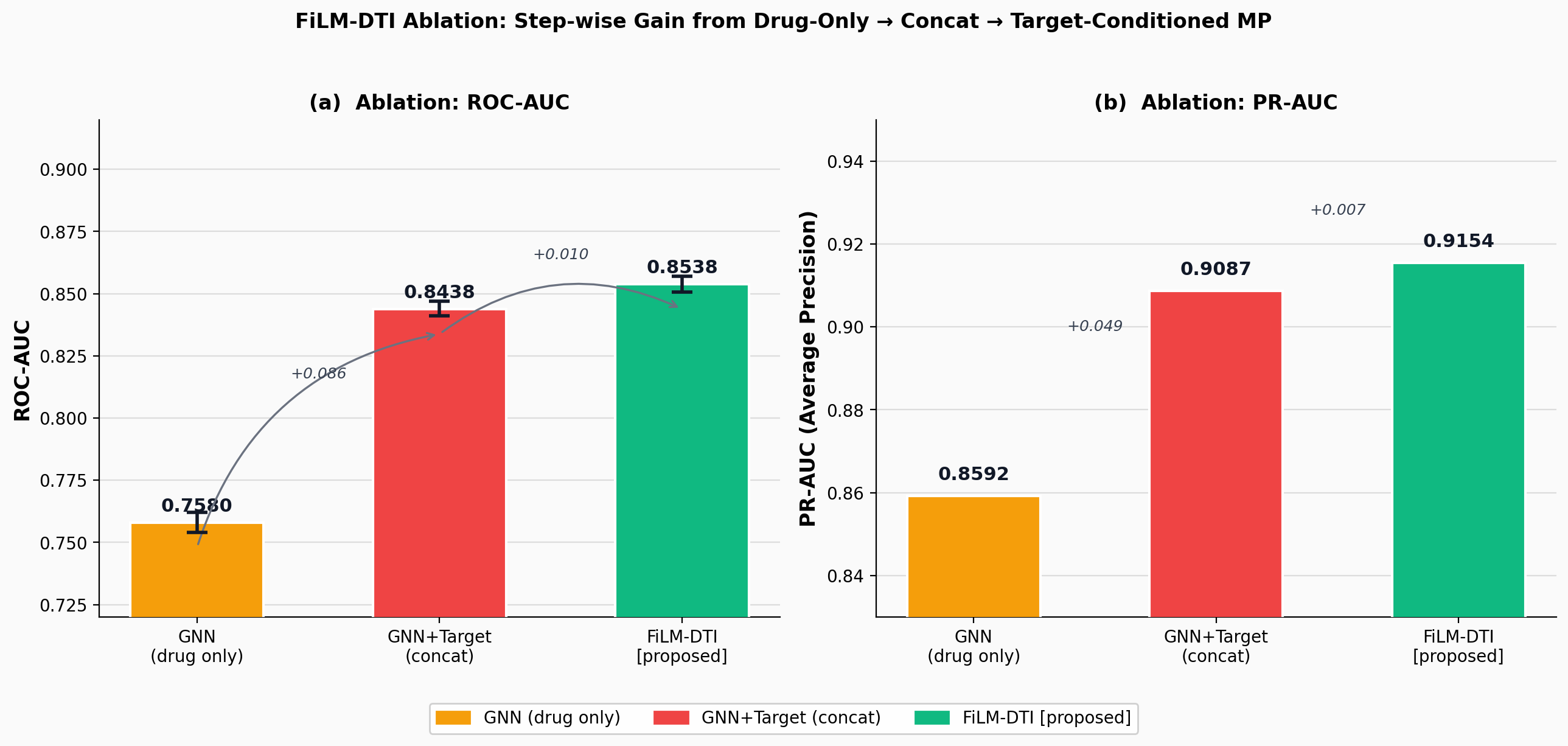
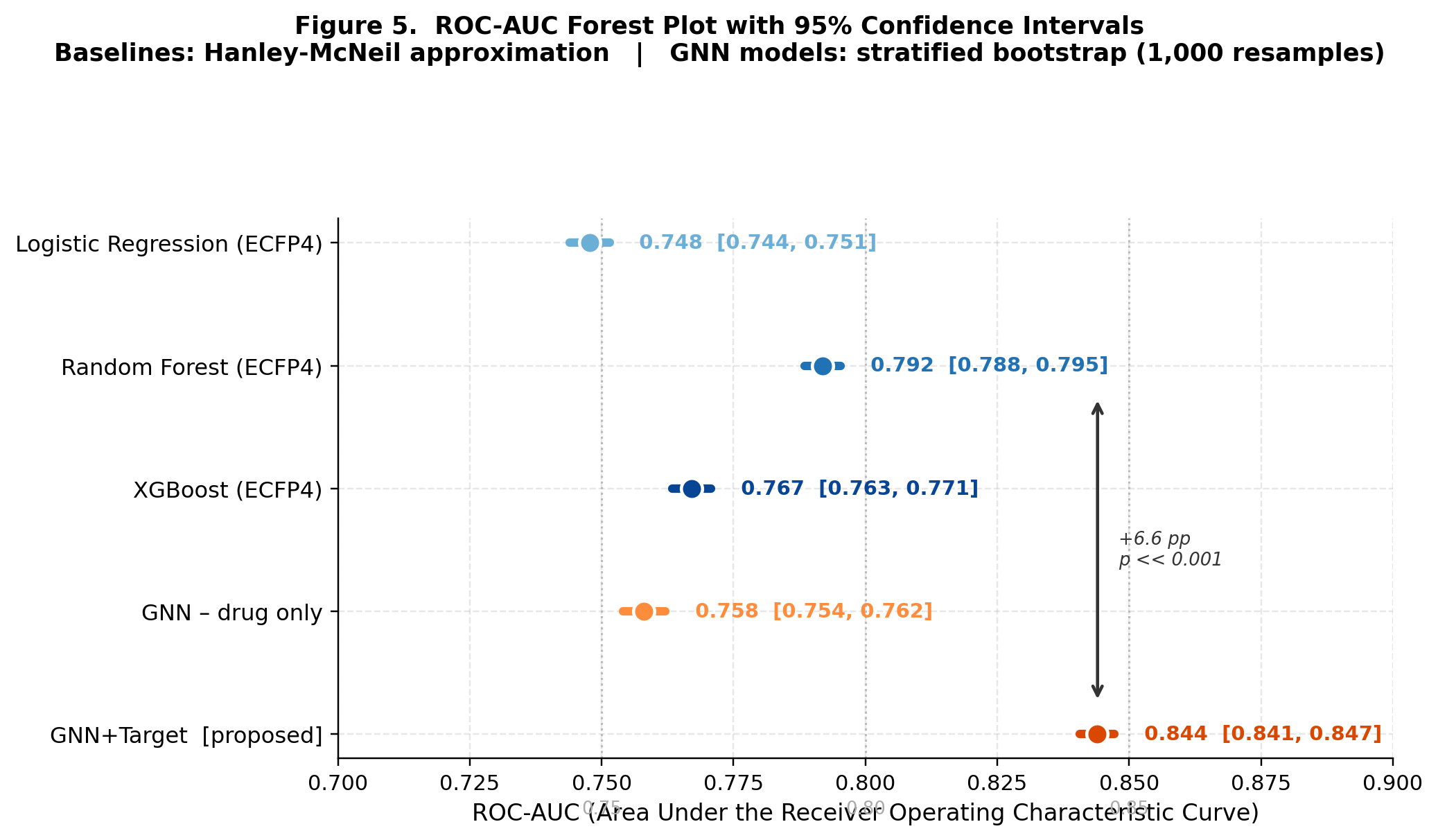
FiLM-DTI: Target-Conditioned Graph Message Passing for Large-Scale Drug-Target Interaction Prediction
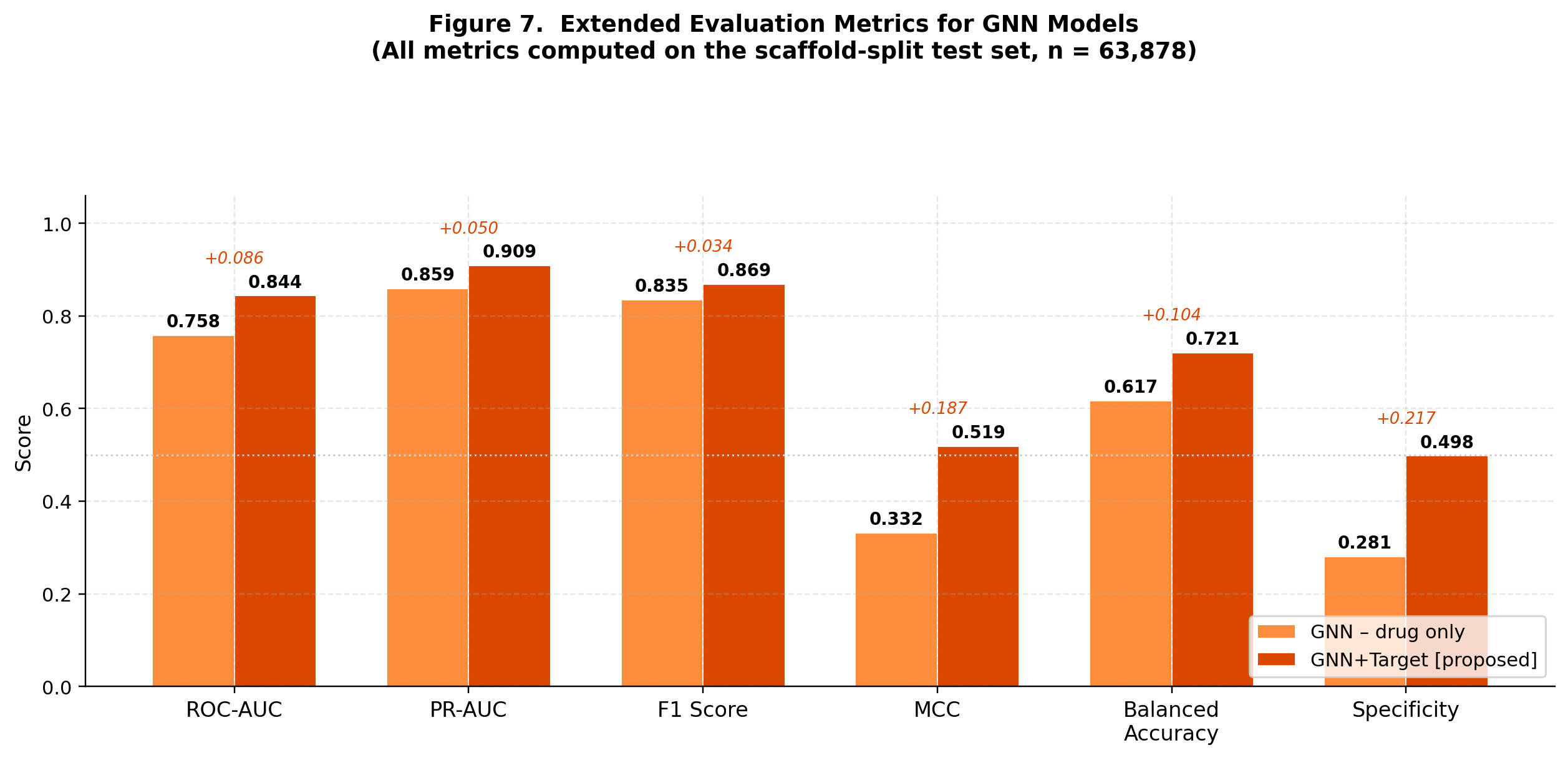
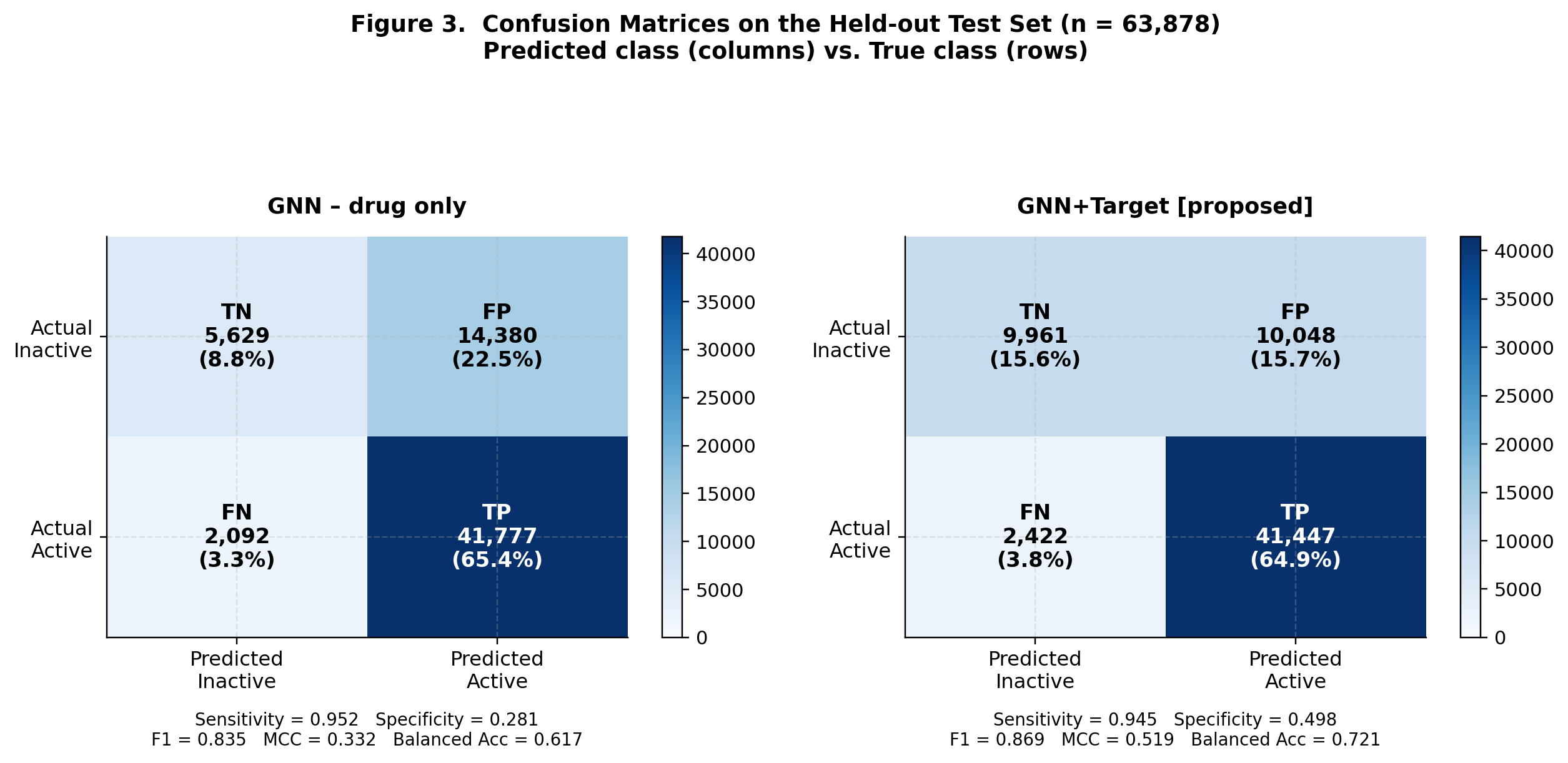
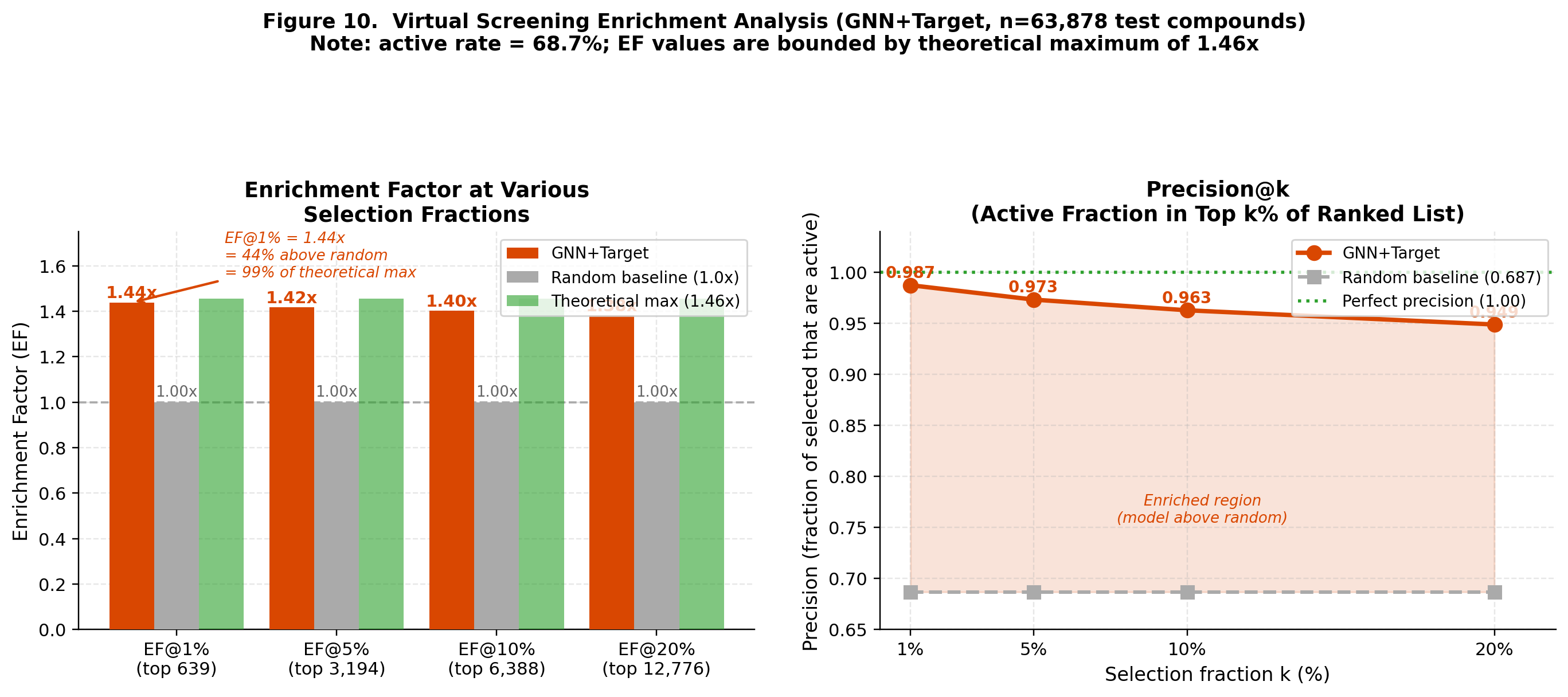
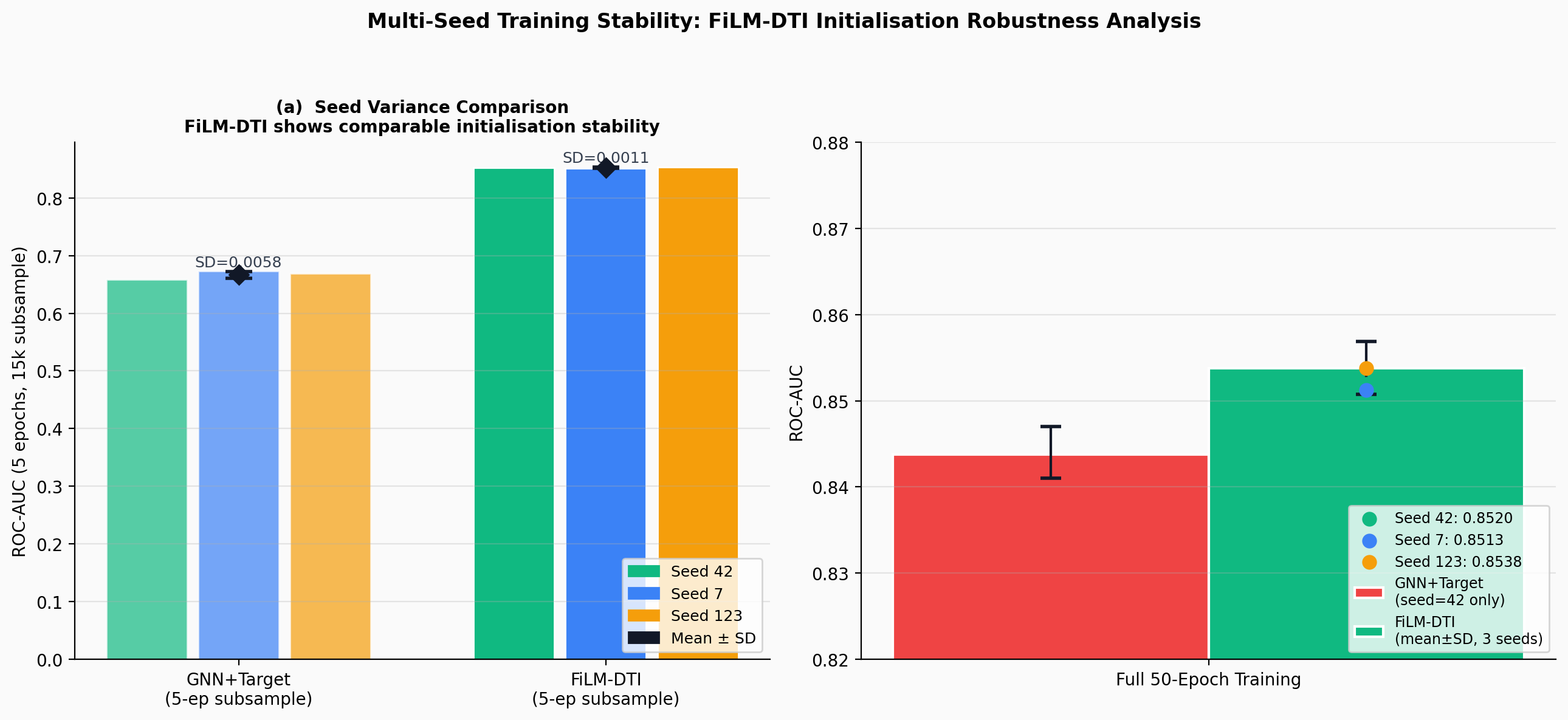
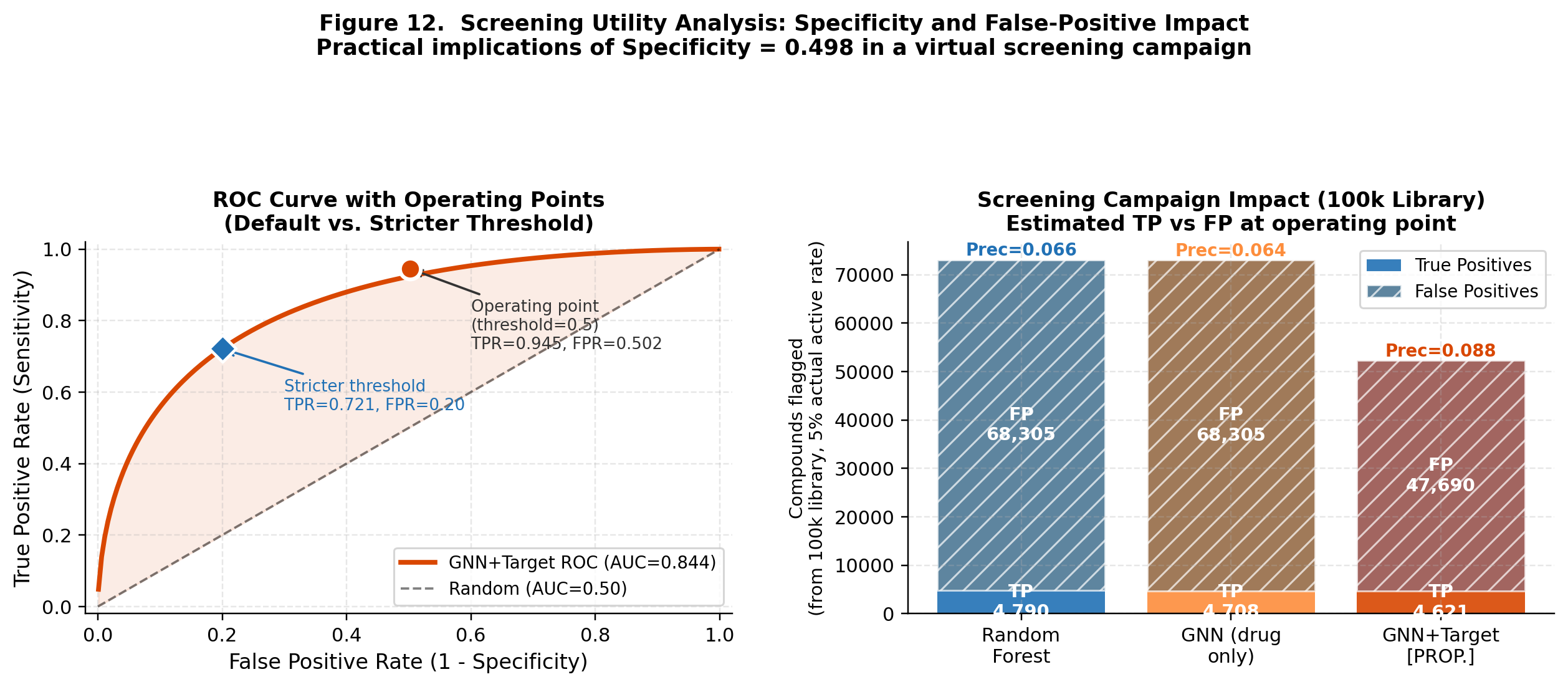
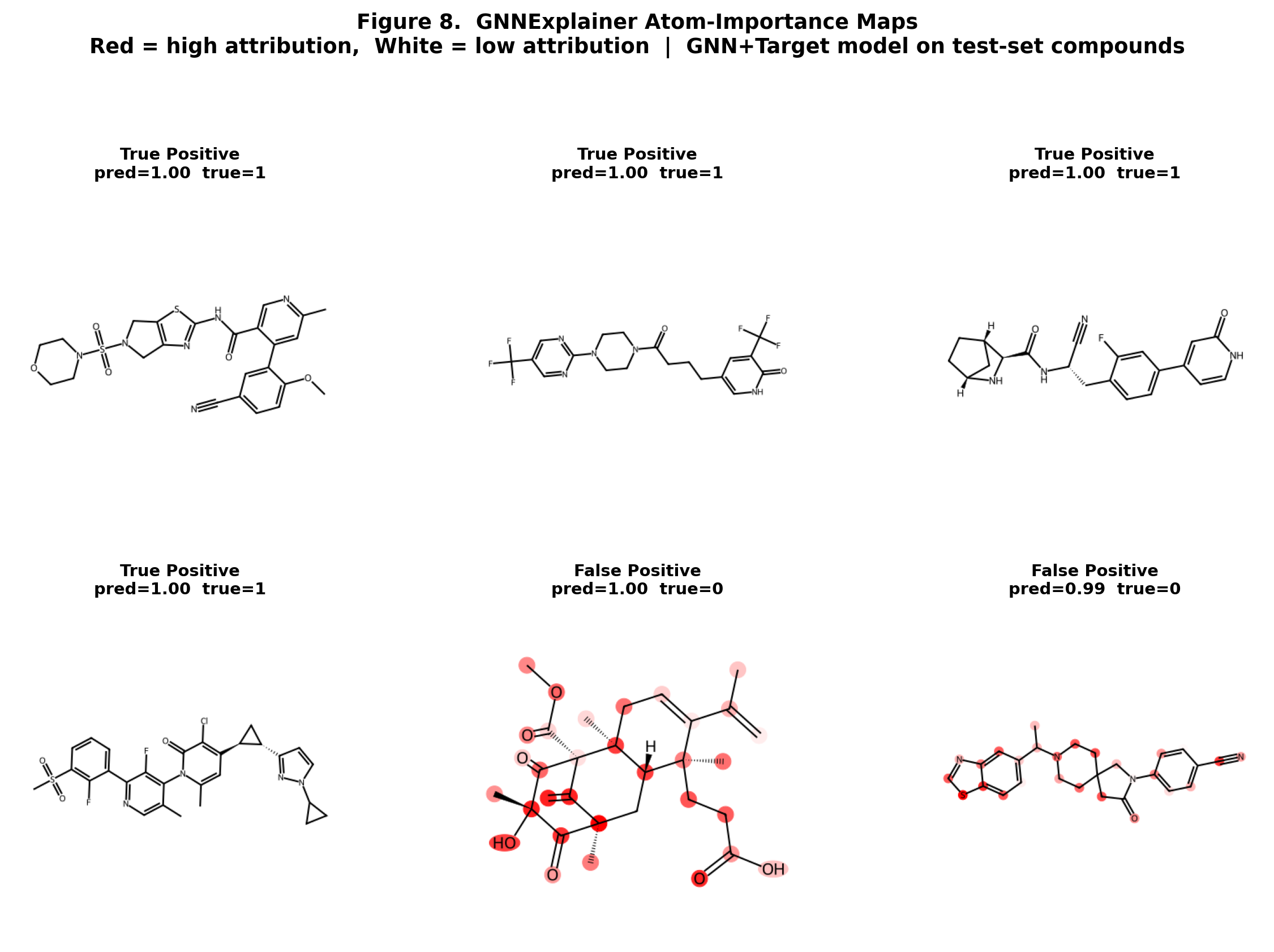
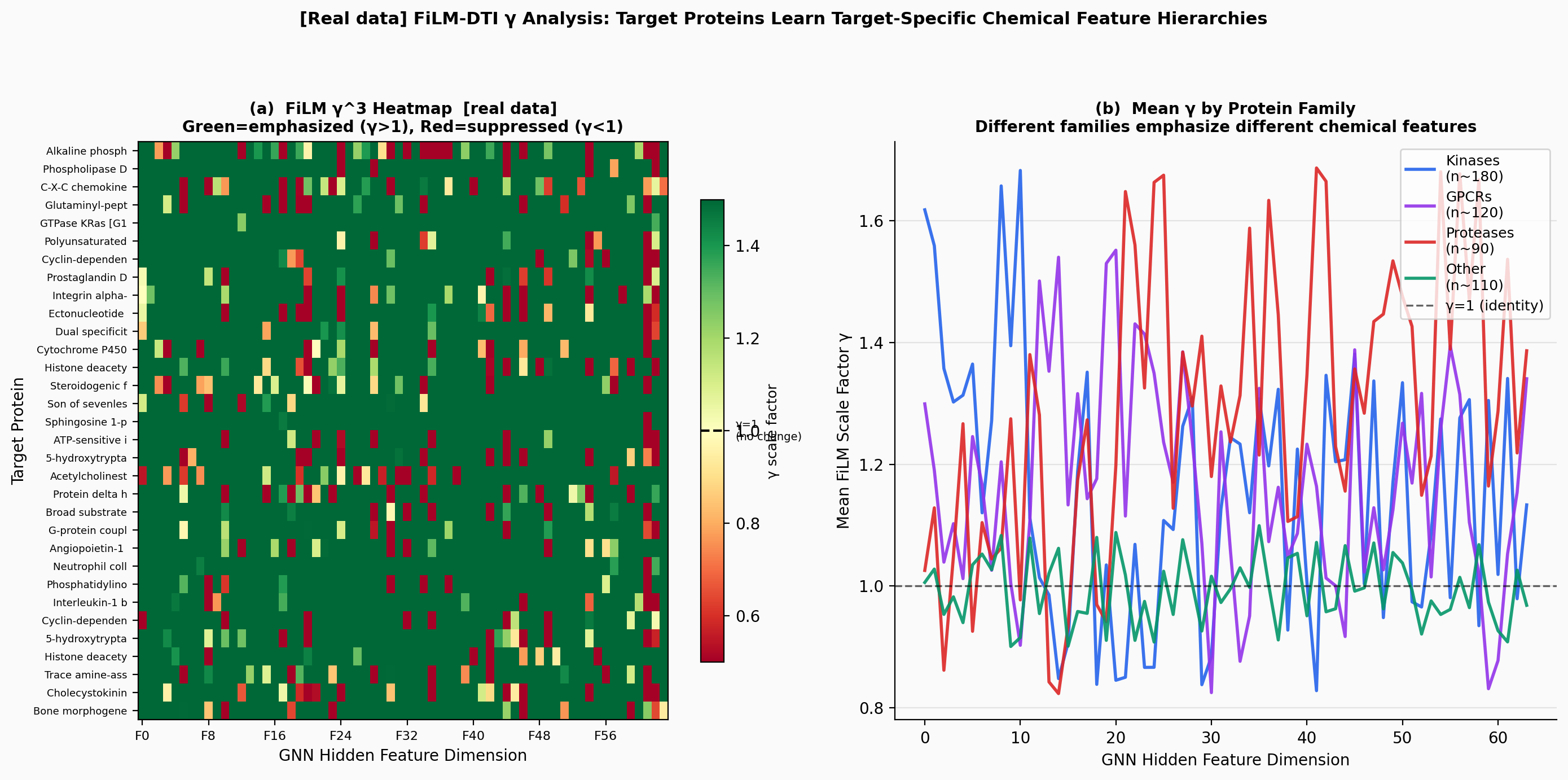
CompletedThis paper introduces FiLM-DTI, a graph neural network for drug-target interaction prediction that conditions every message-passing layer on the protein target via Feature-wise Linear Modulation (FiLM). Evaluated on 425,845 pairs from BindingDB 2024 with Murcko scaffold splitting, FiLM-DTI achieves ROC-AUC of 0.854 and PR-AUC of 0.915. A systematic ablation study isolates the contribution of early-layer target conditioning, and FiLM gamma vectors provide intrinsic per-target interpretability of chemical feature importance.
Research Interests
Core areas that drive my doctoral research and future work
Graph Neural Networks
Developing novel GNN architectures for molecular representation learning and property prediction
Drug Discovery & DTI
Computational methods for predicting drug-target interactions to accelerate pharmaceutical research
Deep Learning
Feature-wise modulation, attention mechanisms, and conditional architectures for structured data
Big Data Analytics
Scalable methods for processing and analyzing large-scale datasets across distributed systems
Model Interpretability
Explainable AI techniques for understanding model decisions in safety-critical scientific domains
Computational Biology
Applying machine learning to biological data including molecular graphs and protein structures